Operator Learning & Neural Operators
From Separation of Variables to Fourier Neural Operators
ML for Science and Engineering · Lecture 18
Joseph Bakarji
Where We Are
A spatio-temporal PDE solution $u(x, t)$, e.g. advection-diffusion
Shortcoming: for a different $u_0(x)$, you must retrain from scratch.
Part I: From Vectors to Functions
What are operators, and why should neural networks learn them?
What Is an Operator?
Maps vectors to vectors:
$f_\theta: \mathbb{R}^n \to \mathbb{R}^m$
Input: $\mathbf{x} = [x_1, \ldots, x_n]$
Output: $\mathbf{y} = [y_1, \ldots, y_m]$
Maps functions to functions:
$\mathcal{G}_\theta: \mathcal{U} \to \mathcal{V}$
Input: $u(\cdot)$ (e.g., initial condition)
Output: $\mathcal{G}(u)(\cdot)$ (e.g., solution)
Example: $u_0(x) \mapsto u(x, T)$: given an initial temperature distribution, predict the temperature at time $T$.
Why Functions, Not Vectors?
A vector $[u_1, u_2, \ldots, u_n]$ is just a discretization of a function $u(x)$ on a grid.
If we think in terms of functions, the learned operator should work at any resolution: coarse or fine.
This is fundamentally different from CNNs, which are tied to a fixed grid resolution.
Universal Approximation for Operators
Chen & Chen (1995): A neural network with a single hidden layer can approximate any continuous nonlinear operator. [paper]
$\psi_i(y)$ , basis functions at the query point (the trunk)
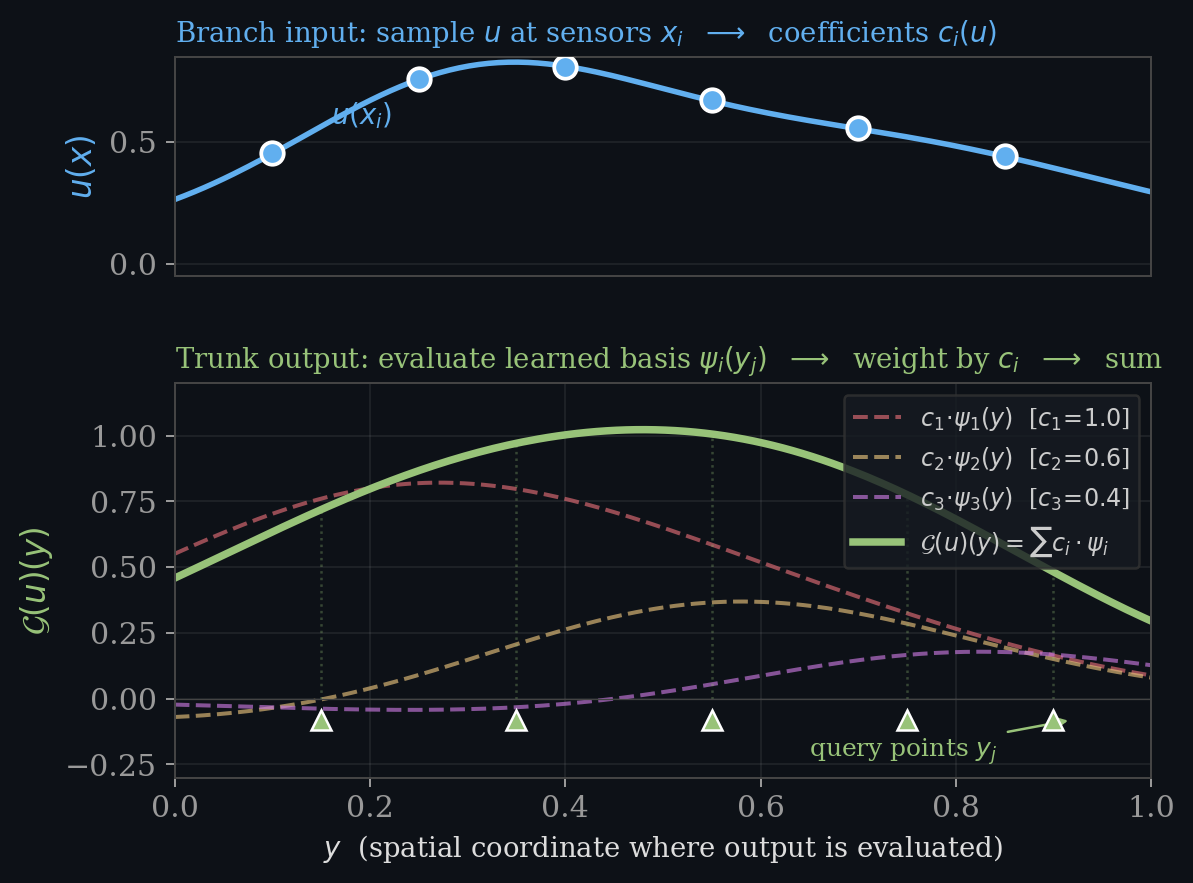
This is a modal decomposition: basis functions $\times$ coefficients, summed. It is the theoretical foundation for DeepONet.
The Operator Learning Setup
$u_0(x)$, BCs, forcing $f$
$\mathcal{G}_\theta$
$u(x,t)$ at any $(x,t)$
Training data: many (input function, solution) pairs from a classical PDE solver.
Once trained: instant evaluation for any new input (no retraining needed).
- Deep Operator Networks (DeepONet) , inspired by Chen & Chen's universal approximation theorem [Lu et al. 2021]
- Fourier Neural Operators , inspired by Green's functions and Fourier transformations [Li et al. 2021]
Part II: Deep Operator Networks
From classical modal decomposition to learned operators.
Spatial-Temporal Modal Decomposition
Many PDE solutions can be written as a sum of spatial modes weighted by temporal coefficients:
You have seen this in multiple forms:
- SVD/POD (Lecture 9): orthogonal modes from data ($U \Sigma V^\top$)
- DMD (Lecture 10): dynamic modes with eigenvalue evolution
- Fourier series: sines and cosines as basis functions
DeepONet Architecture
Two sub-networks whose outputs are combined via a dot product (Lu et al., Nature Machine Intelligence, 2021): [paper]
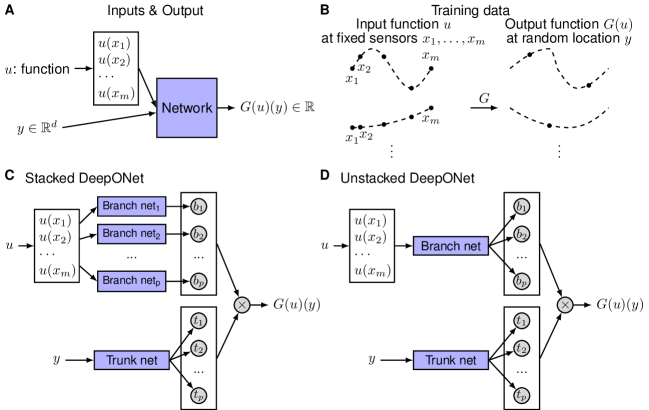
DeepONet = Learned Modal Decomposition
$u(x,t) \approx \sum_k a_k \cdot \varphi_k(x)$
$\varphi_k(x)$: eigenvectors (fixed)
$a_k$: projection coefficients (linear)
$\mathcal{G}(u)(y) \approx \sum_i b_i(u) \cdot t_i(y)$
$t_i(y)$: trunk outputs (learned basis)
$b_i(u)$: branch outputs (nonlinear in $u$)
Training DeepONet
Supervised training: given $N$ (input function, solution) pairs from a PDE solver:
Here $u_k$ is the $k$-th initial condition (sampled at sensor locations), and $y_j$ are query points where we evaluate the output solution.
$\bullet$ sensors = where $u_k$ is sampled • $\bullet$ $y_j$ = query points
Physics-informed (PI-DeepONet): add PDE residual loss where $f$ is the forcing/source term:
DeepONet: Key Properties
| Property | DeepONet |
|---|---|
| Irregular geometries | ✓ trunk evaluates at any point |
| Variable sensor locations | ✓ branch takes arbitrary inputs |
| Noise robustness | ✓ error increases <10× at 0.1% noise |
| No grid requirement | ✓ fully meshless |
| Resolution invariance | ~ approximate (depends on sensor density) |
DeepONet: Results
From Lu et al. (2021), DeepONet achieves high accuracy across diverse PDE families:
| Problem | Relative $L^2$ Error | Notes |
|---|---|---|
| Advection equation | $\sim\!10^{-3}$ | Linear, smooth solutions |
| Diffusion-reaction | $\sim\!10^{-3}$ | Nonlinear source term |
| Stochastic ODE system | $\sim\!10^{-2}$ | Random forcing |
| Gravity pendulum | $\sim\!10^{-3}$ | Nonlinear ODE |
Lu et al. (2021). Learning nonlinear operators via DeepONet. Nature Machine Intelligence, 3, 218–229. [link]
Part III: Green's Functions → Fourier Neural Operators
From integral solutions to learnable spectral layers.
Green's Functions: The Integral Solution
The Green's function $G(x,y)$ is the solution to the PDE when the forcing is a single impulse at $y$:
It tells you how a point source at $y$ influences the solution at $x$. Because the PDE is linear, the full solution is a superposition of these impulse responses:
Superposition: Interactive
For the 1D diffusion equation $\;\frac{\partial u}{\partial t} = \nu \frac{\partial^2 u}{\partial x^2} + f(x)$, the Green's function is a Gaussian kernel. Add impulses to $f(x)$ and see the superposition build the solution $u(x)$.
From Green's Functions to Neural Operator Layers
Green's functions rely on linearity (superposition). For nonlinear PDEs, no simple $G$ exists. But we can learn a kernel and compose it through nonlinear layers.
$W$ is a finite matrix (fixed input dimension)
$\kappa(x,y)$ is a learnable kernel (continuous, any resolution)
Kovachki, N. et al. (2023). Neural Operator: Learning Maps Between Function Spaces. JMLR, 24(89), 1–97.
The Convolution Trick
A general kernel $\kappa(x, y)$ is expensive: $O(n^2)$ to evaluate on a grid of $n$ points.
Assume translation invariance: the kernel depends only on the difference $\kappa(x, y) = \kappa(x - y)$.
→ This is convolution!
→ Convolution in space = multiplication in Fourier domain.
FNO Layer: Interactive
Each Fourier Neural Operator layer: apply FFT, multiply by learned weights, inverse FFT, add spatial path, activate.
FNO: Key Properties
| Property | FNO |
|---|---|
| Resolution invariance | ✓ same parameters at any grid size |
| Computational cost | ✓ $O(n \log n)$ per layer (via FFT) |
| Regular grids only | ✗ requires uniform Cartesian mesh |
| Spectral bias | ✗ mode truncation discards high frequencies |
| Speed vs. classical | ✓ 1000× faster than pseudo-spectral solvers |
Li, Z., Kovachki, N., Azizzadenesheli, K., Liu, B., Bhatt, K., Stuart, A. & Anandkumar, A. (2021). Fourier Neural Operator for Parametric PDEs. ICLR 2021.
In Code: DeepONet
class DeepONet(nn.Module):
def __init__(self, u_dim, y_dim, p=64):
super().__init__()
self.branch = nn.Sequential( # input function → coefficients
nn.Linear(u_dim, 128), nn.ReLU(),
nn.Linear(128, p))
self.trunk = nn.Sequential( # query point → basis functions
nn.Linear(y_dim, 128), nn.ReLU(),
nn.Linear(128, p))
def forward(self, u, y):
b = self.branch(u) # (batch, p)
t = self.trunk(y) # (batch, p)
return (b * t).sum(-1) # dot productIn Code: FNO Layer
class SpectralConv1d(nn.Module):
def __init__(self, ch, modes):
super().__init__()
self.modes = modes
self.weights = nn.Parameter( # learnable Fourier weights
torch.randn(ch, ch, modes, 2))
def forward(self, x): # x: (batch, ch, N)
x_ft = torch.fft.rfft(x)
out_ft = torch.zeros_like(x_ft)
out_ft[:,:,:self.modes] = \ # multiply low modes by R
complex_mul(x_ft[:,:,:self.modes], self.weights)
return torch.fft.irfft(out_ft, n=x.size(-1))Operator Learning: DeepONet & FNO Hands-On
Operator Learning: DeepONet & FNO Hands-On
In this notebook we implement minimal versions of DeepONet and Fourier Neural Operator (FNO) and train them on a 1D diffusion equation.
Problem Setup
We learn the solution operator for the 1D heat equation:
$$\frac{\partial u}{\partial t} = \nu \frac{\partial^2 u}{\partial x^2}, \quad u(x, 0) = u_0(x)$$
Given random initial conditions $u_0(x)$, the operator maps $u_0 \mapsto u(x, T)$ at a fixed time $T$.
Data Generation
We generate random initial conditions as sums of Fourier modes with decaying amplitudes, then solve the heat equation exactly using the spectral method (FFT-based). This gives us ground-truth (input, output) pairs for training.
Part 1: DeepONet
The branch network takes the input function $u_0$ (sampled at sensor locations) and produces coefficients $b_1, \ldots, b_p$.
The trunk network takes query points $y$ and produces basis functions $t_1(y), \ldots, t_p(y)$.
$$\mathcal{G}(u_0)(y) = \sum_{i=1}^{p} b_i(u_0) \cdot t_i(y) = \langle \mathbf{b}(u_0), \mathbf{t}(y) \rangle$$
DeepONet Architecture
The model below has two sub-networks:
- Branch (
nn.Sequential): takes the full input function $u_0$ sampled at $N$ sensor points and outputs $p$ coefficients. - Trunk (
nn.Sequential): takes query coordinates $y$ and outputs $p$ basis function values.
The output is their dot product: $\mathcal{G}(u_0)(y) = \langle \mathbf{b}(u_0), \mathbf{t}(y) \rangle + \text{bias}$.
Part 2: Fourier Neural Operator (FNO)
Each FNO layer applies: $v^{(\ell+1)} = \sigma\big(W v^{(\ell)} + \mathcal{F}^{-1}(R_\phi \cdot \mathcal{F}(v^{(\ell)}))\big)$
- FFT the input, multiply low modes by learned weights $R_\phi$, iFFT back
- Add a local linear transform $W v$ (skip connection in spatial domain)
- Apply nonlinear activation
FNO Architecture
The key building block is SpectralConv1d: it applies the FFT, multiplies the lowest modes Fourier coefficients by a learned complex weight matrix $R_\phi$, then applies the inverse FFT.
The full FNO1d model stacks multiple such layers. Each layer also has a local 1x1 convolution $W$ (skip connection in spatial domain). The input is first "lifted" from 1 channel to width channels, and the output is projected back to 1 channel.
Comparison
Let's compare DeepONet and FNO side by side on the same test samples.
Exercises
-
Resolution transfer (FNO): Train the FNO on a 64-point grid, then evaluate it on a 128-point grid. Does it transfer? Why?
-
Sensor sparsity (DeepONet): Reduce the number of sensor points in the branch network (e.g., use every 4th point). How does accuracy degrade?
-
Harder PDE: Replace the heat equation with Burgers' equation $u_t + u u_x = \nu u_{xx}$. Which method handles the nonlinearity better?
-
Physics-informed DeepONet: Add a PDE residual loss term $|\mathcal{N}[\mathcal{G}_\theta(u_0)] - 0|^2$ and train with fewer data pairs. Does it help?
Part IV: Comparing Architectures
DeepONet vs FNO vs PINNs: when to use what.
When to Use What
| Feature | DeepONet | FNO | PINNs |
|---|---|---|---|
| Irregular geometry | ✓✓ | ✗ | ✓ |
| Resolution invariance | ~ | ✓✓ | ✗ |
| Noise robustness | ✓✓ | ✗ | ~ |
| No training data needed | ✗ | ✗ | ✓ |
| Generalize over ICs | ✓ | ✓ | ✗ |
| Speed (inference) | ✓ ~ms | ✓✓ ~ms | ✗ retrain |
Other Operator Architectures
The field is evolving rapidly. Some notable variants:
Replace Fourier basis with POD modes learned from data. Better for problems with dominant coherent structures.
Kernel as message-passing on graphs. Works on irregular meshes. $O(n^2)$ complexity.
Attention-weighted averaging of features. Strong with sparse measurements. (Kissas et al., JMLR 2022)
No single architecture dominates. Choice depends on geometry, data quality, and problem structure.
Connections Across the Course
→ DeepONet trunk learns generalized basis functions
→ Temporal dynamics of modes, eigenvalue evolution
→ PI-DeepONet adds PDE loss to operator training
→ Branch-trunk split is a neural generalization
Summary: Design Principles
Operators map functions to functions, generalizing beyond fixed-size vectors. Discretization invariance is the goal.
Modal decomposition → DeepONet.
Green's function → FNO.
Inductive bias from mathematics.
The key advantage over PINNs: learn the operator once, then instantly solve for any new input.
References
Chen, T. & Chen, H. (1995). Universal approximation to nonlinear operators by neural networks with arbitrary activation functions. IEEE Trans. Neural Networks, 6(4), 911–917.
Lu, L., Jin, P., Pang, G., Zhang, Z. & Karniadakis, G.E. (2021). Learning nonlinear operators via DeepONet. Nature Machine Intelligence, 3, 218–229.
Li, Z., Kovachki, N., Azizzadenesheli, K. et al. (2021). Fourier Neural Operator for Parametric PDEs. ICLR 2021.
Kovachki, N., Li, Z., Liu, B. et al. (2023). Neural Operator: Learning Maps Between Function Spaces. JMLR, 24(89), 1–97.